CORE CAPABILITY
Sequence Workspace
Edit sequences with the context you need—features, overlays, and annotations tied to the underlying sequence. Design primers, plan digests, and stage cloning workflows without losing provenance.
PRODUCT VIEW
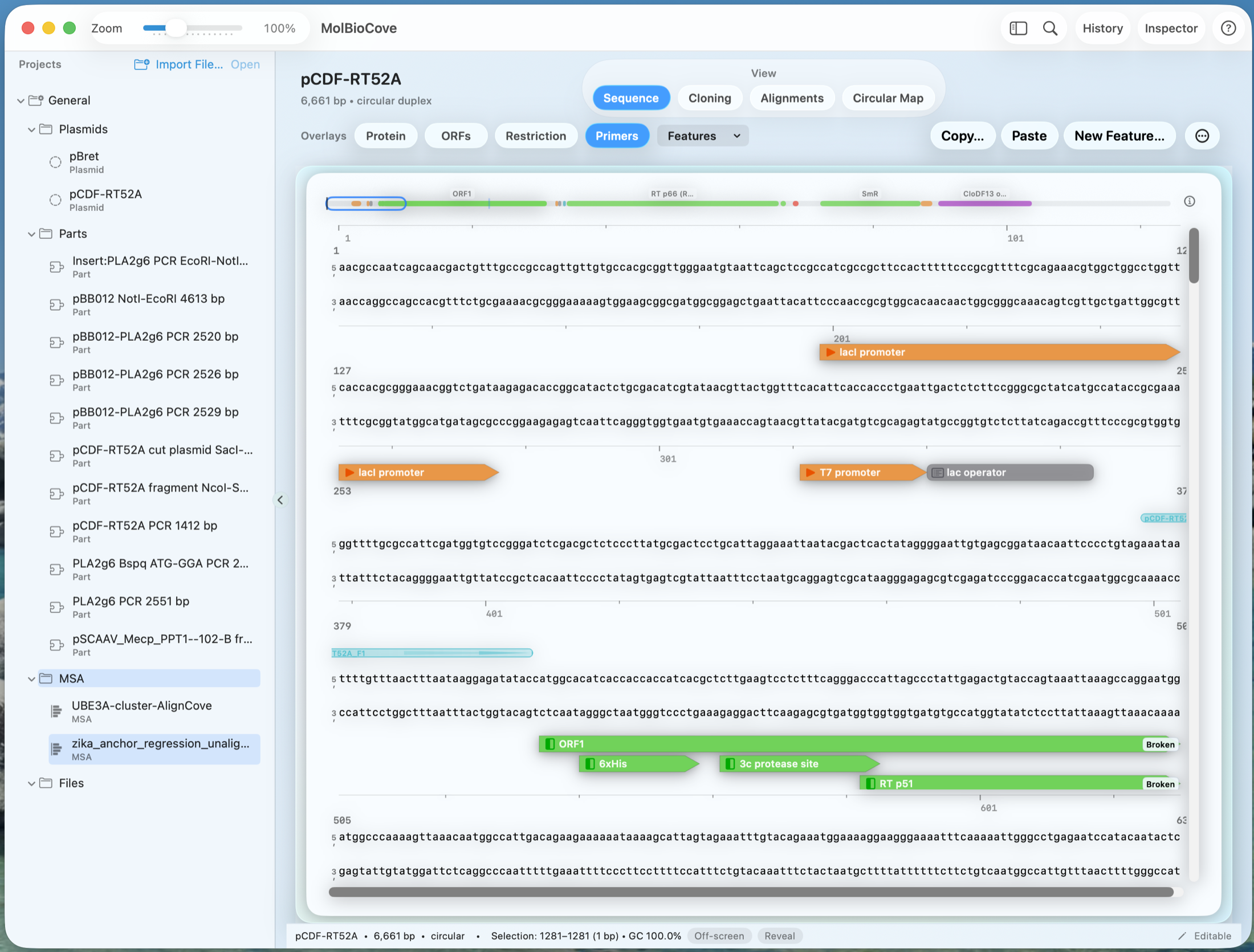
Sequence Workspace in action
Overlays (Protein, ORFs, Restriction, Primers) stay tied to the sequence—so you immediately see what a change affects.

WORKSPACE TOOLS
Primers, digests, and cloning—without leaving the sequence
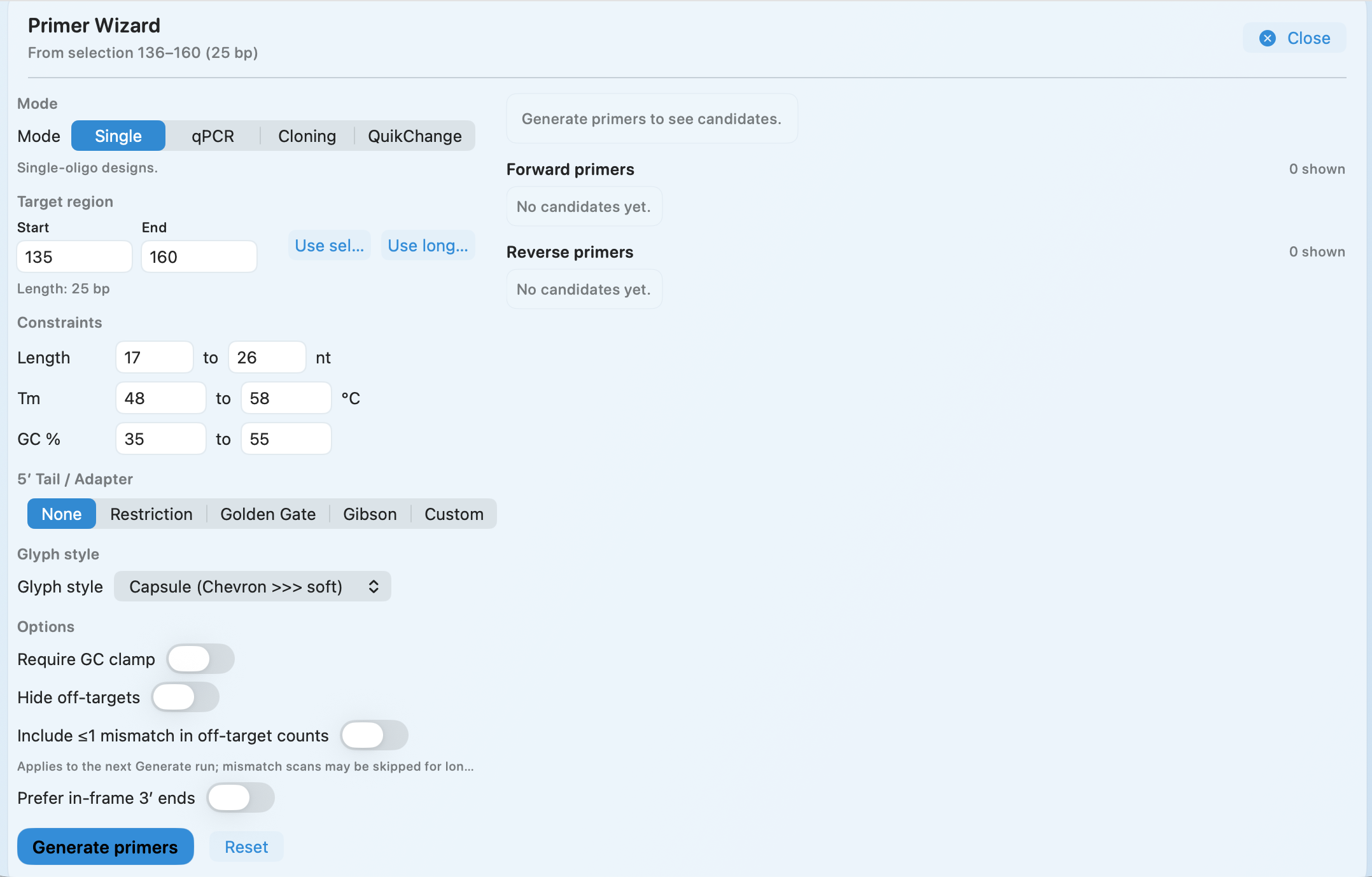
Primer Wizard
Generate primers from any selection with constraints you control—and keep them tied to the exact sequence context they were designed against.
- Single, qPCR, cloning, and QuikChange-style mutagenesis modes
- Tune length, Tm, and GC% with practical guardrails (including a GC clamp)
- Optional off-target checks and mismatch handling
- Add 5′ tails for restriction sites, Golden Gate, Gibson, or custom adapters
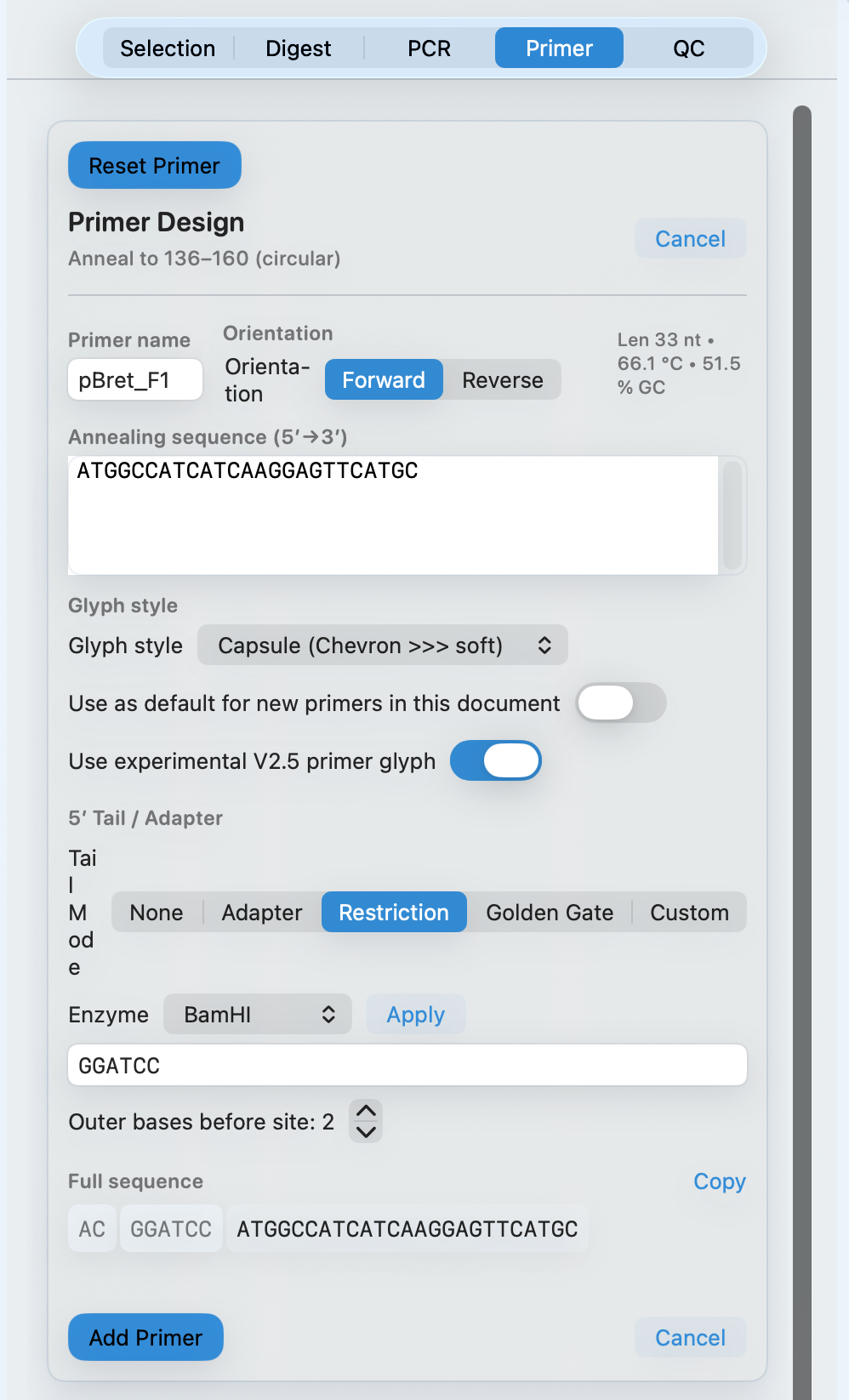
Add restriction sites in one click
When you already know the enzyme, MolBioCove adds the site plus 5′ flanking bases and shows an order-ready sequence.
- Choose an enzyme and set 5′ flanking bases before the site
- See length, Tm, and GC% update live as you edit
- Copy a clean 5′→3′ sequence for ordering
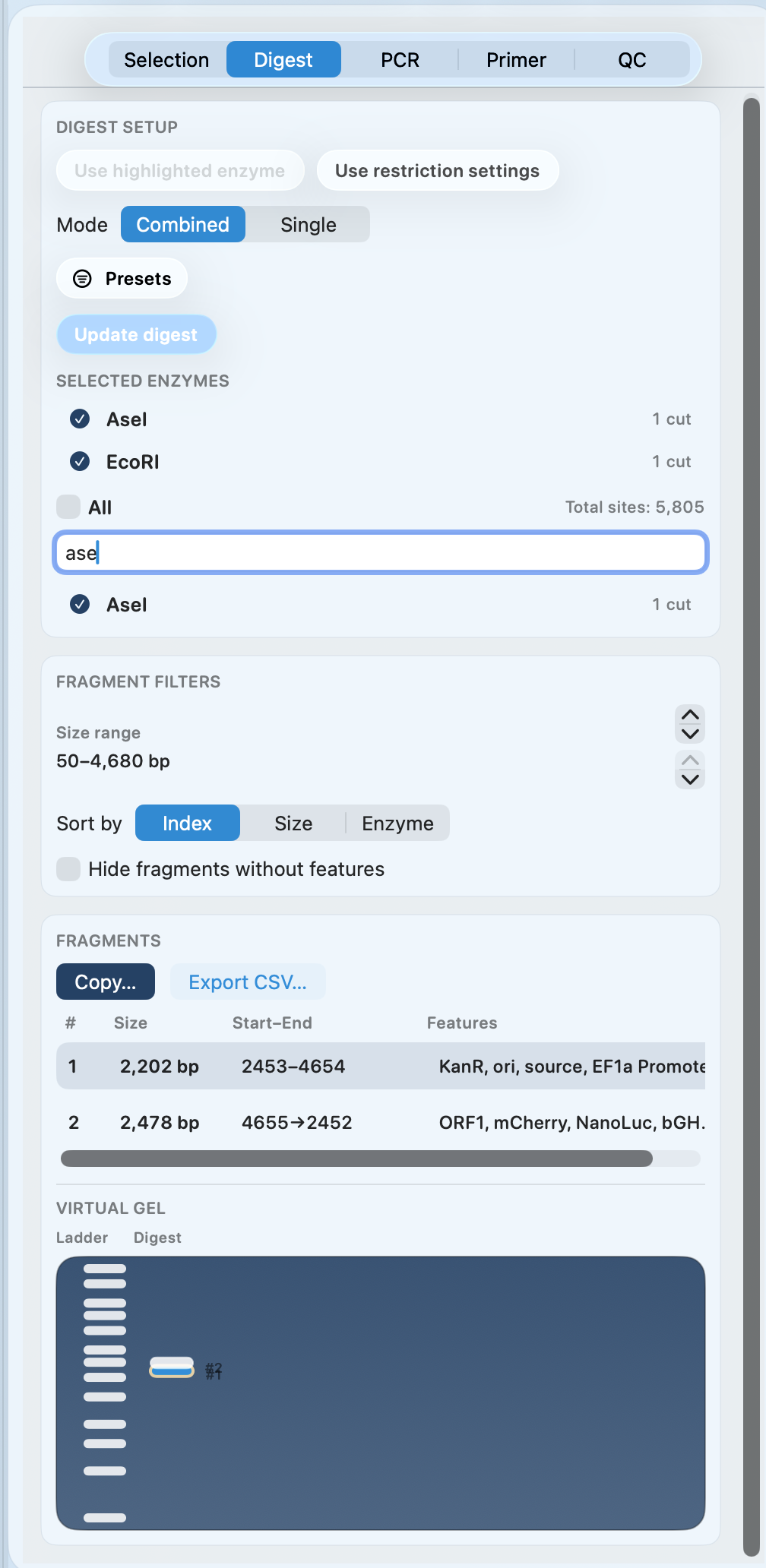
Digest planning with virtual gel preview
Choose enzymes, inspect predicted fragments, and preview a virtual gel—so you pick the right backbone and inserts before you cut DNA.
- Combined and single-enzyme digest modes
- Fragment table with start/end coordinates and feature context
- Copy fragments or export CSV
- Virtual gel preview for a quick sanity check
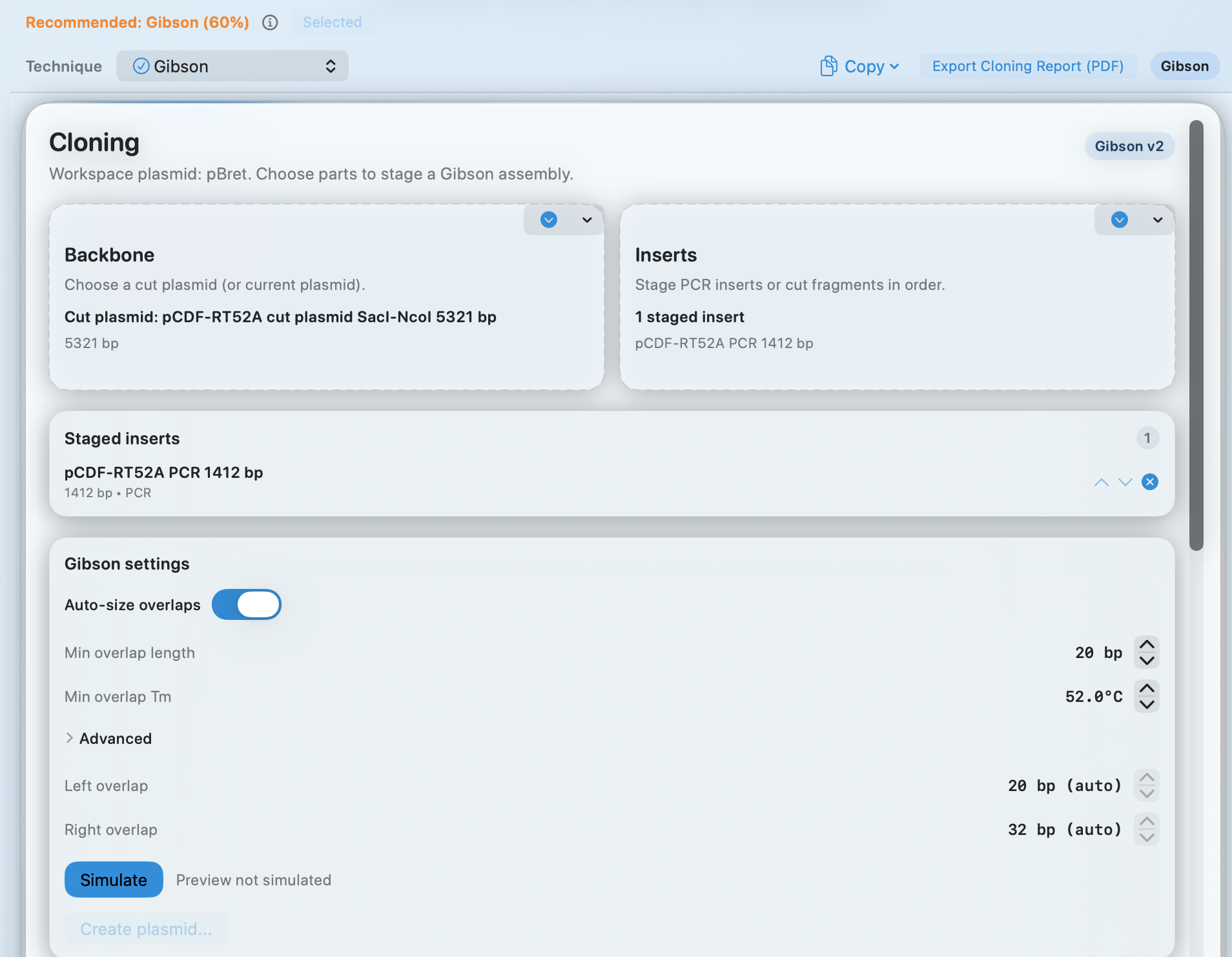
Cloning wizard + simulation
Stage a backbone and inserts, simulate the assembly, and export a report for your notebook—or a clean handoff.
- Technique recommendation (e.g., Gibson) with a confidence hint
- Auto-sized overlaps plus minimum overlap length and Tm controls
- Simulate before committing to a build
- Export a cloning report (PDF)
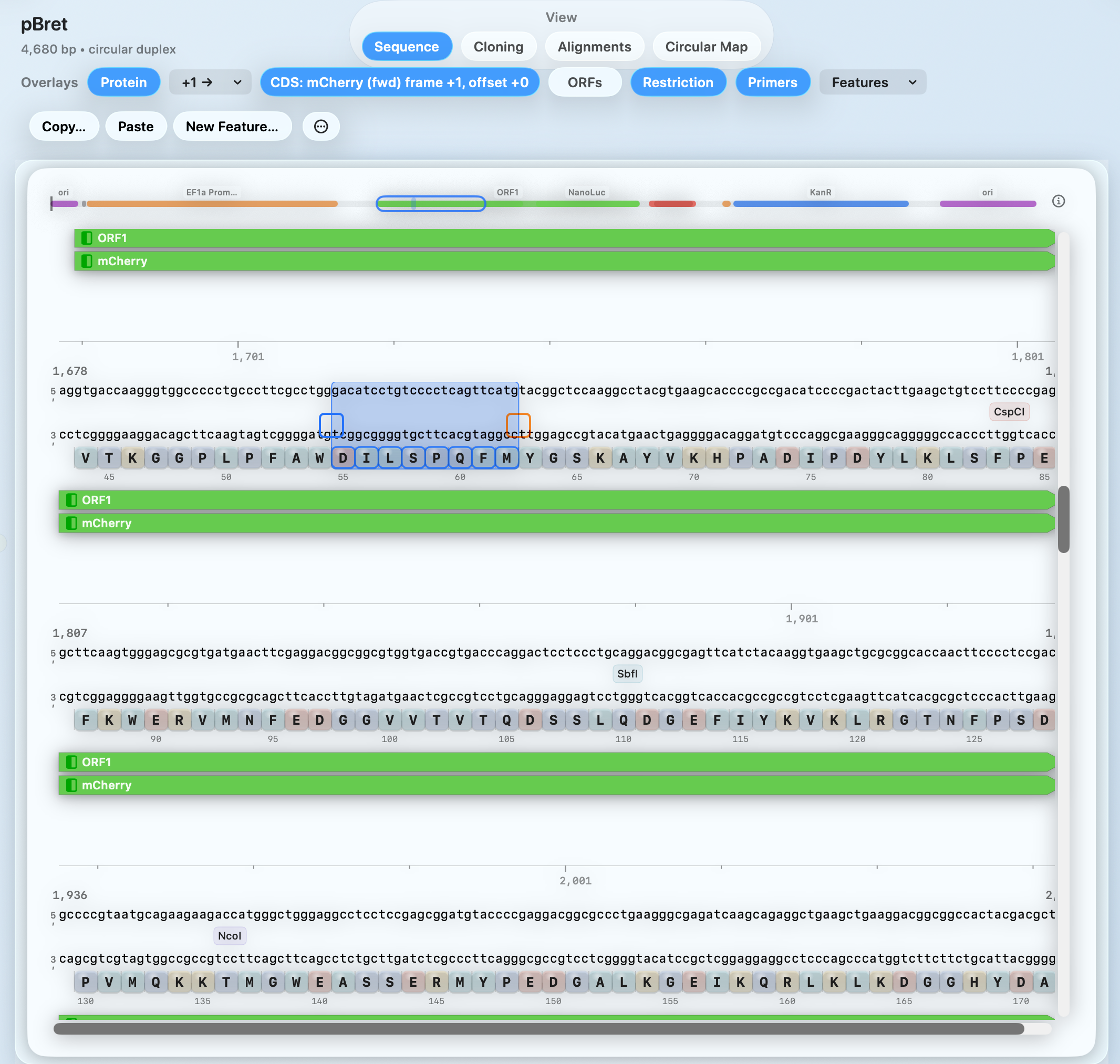
Protein & ORF context while you edit
Translate in-frame, keep amino-acid context visible, and tie AA/codon indexing to the underlying sequence—so protein edits stay deliberate.
- In-frame amino-acid track under the sequence
- Clear AA/codon indexing on hover and selection
- Overlays for ORFs, restriction sites, primers, and features
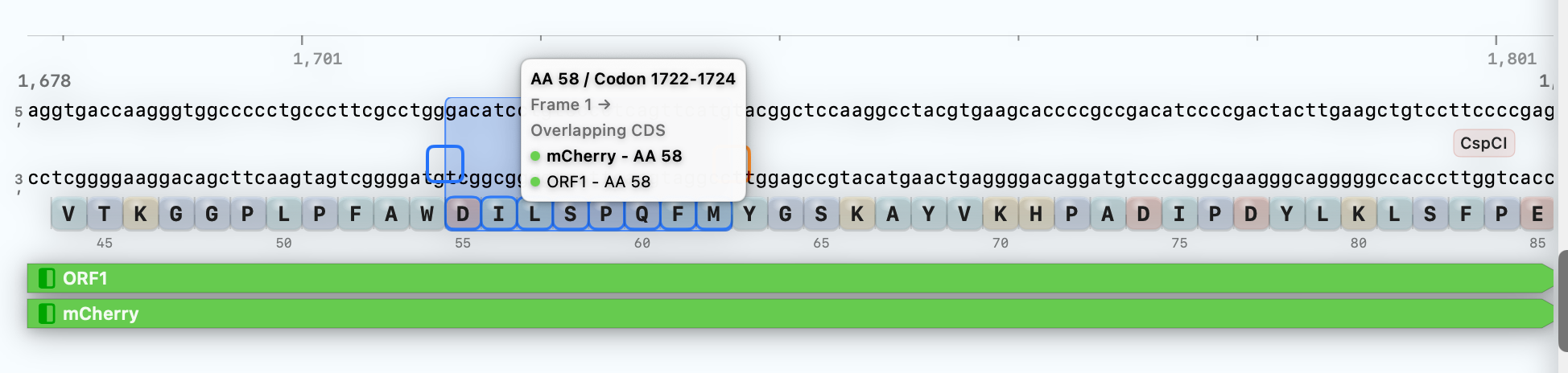
Hover any selection to see amino-acid and codon indexing.
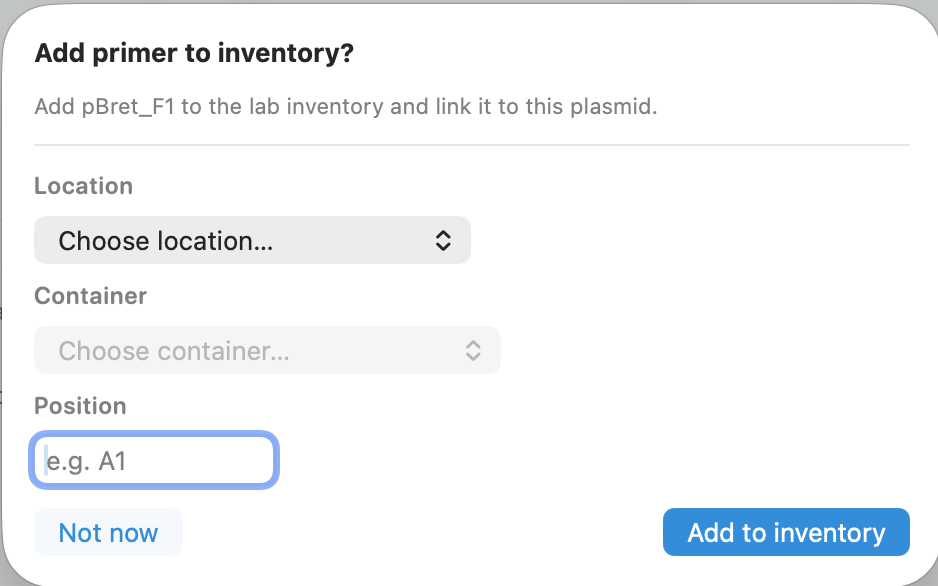
Link primers to physical inventory
Link primers (and plasmids) to where they live—so the right tube is always findable.
- Store location, container, and position (e.g., A1)
- Optional prompt right after primer creation
- Cleaner handoffs and fewer “which tube is this?” moments
WHAT YOU CAN DO
Common tasks
TYPICAL WORKFLOW
A typical flow
Open a construct and bring sequence context, features, and overlays into view.
Make edits while keeping annotations and protein/ORF context visible.
Design primers, plan digests, and sanity-check cloning steps before you commit to a build.
Export primer sequences, fragment lists, and a cloning report—so handoffs stay reviewable.
OUTPUTS
Outputs
Request Beta Access
Join the external beta—priority given to active bench teams.