Built by bench scientists, for bench scientists.
Stop switching tools for construct work.
RayCrest keeps plasmid editing, sequencing review, and publication-ready maps attached to
the same construct in one local-first Mac workspace.
Edit plasmids and build publication-ready plasmid maps
Design primers and plan in silico PCR workflows
Open AB1 files, review chromatograms, and check FASTQ evidence
Run repeatable protein multiple sequence alignment in one alignment workspace
Verified compatibility
GenBank
FASTA
SnapGene .dna import
Sanger .ab1 / .abi
FASTQ
Open GenBank files, FASTA files, SnapGene .dna files, AB1 files, and FASTQ files in the same local workflow.
See interoperability details
POPULAR WORKFLOWS
Five workflows where RayCrest is strongest
The core jobs bench teams reach for most in RayCrest.
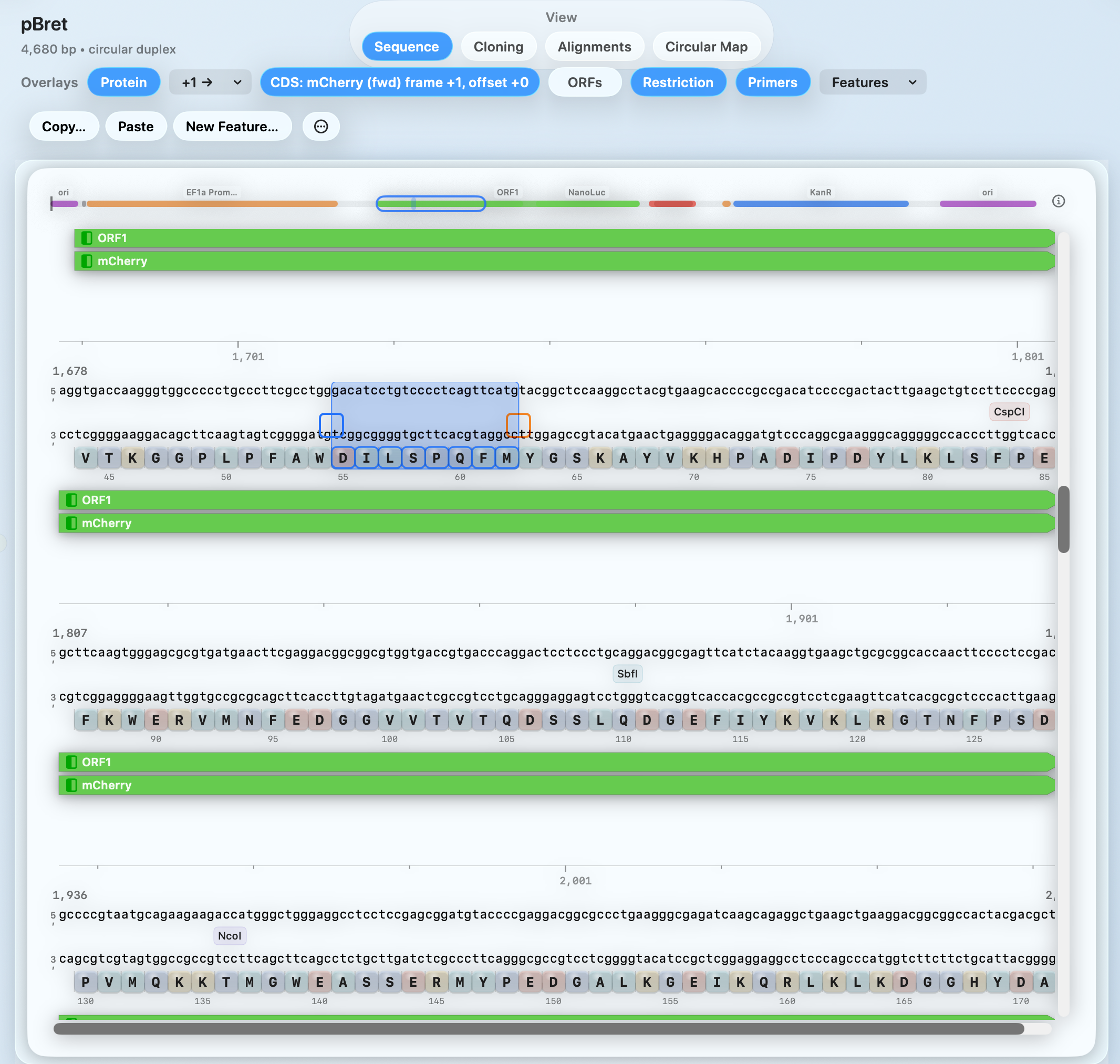
Plasmid Editor
Edit constructs and export publication-ready plasmid maps from the same sequence
Move from construct changes to shareable plasmid maps without rebuilding it in a second tool.
Linked overlays for features, ORFs, restriction sites, proteins, and primers.
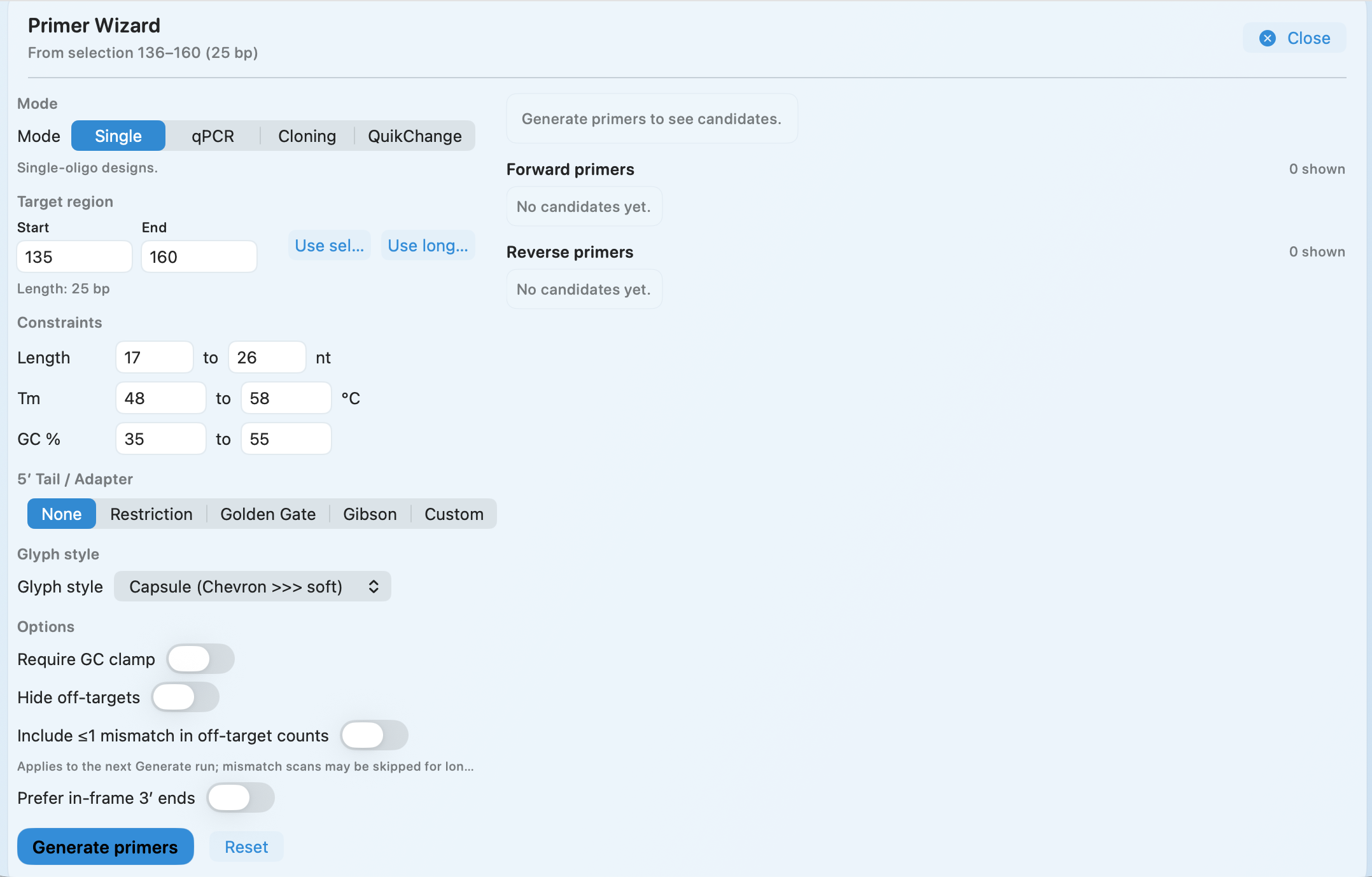
Primer Design
Design primers from the same view as the active construct
Design primers and plan PCR simulation in the same construct view, with modes for sequencing, cloning, and mutagenesis.
Single, qPCR, cloning, and QuikChange-style modes with 5′ tails.
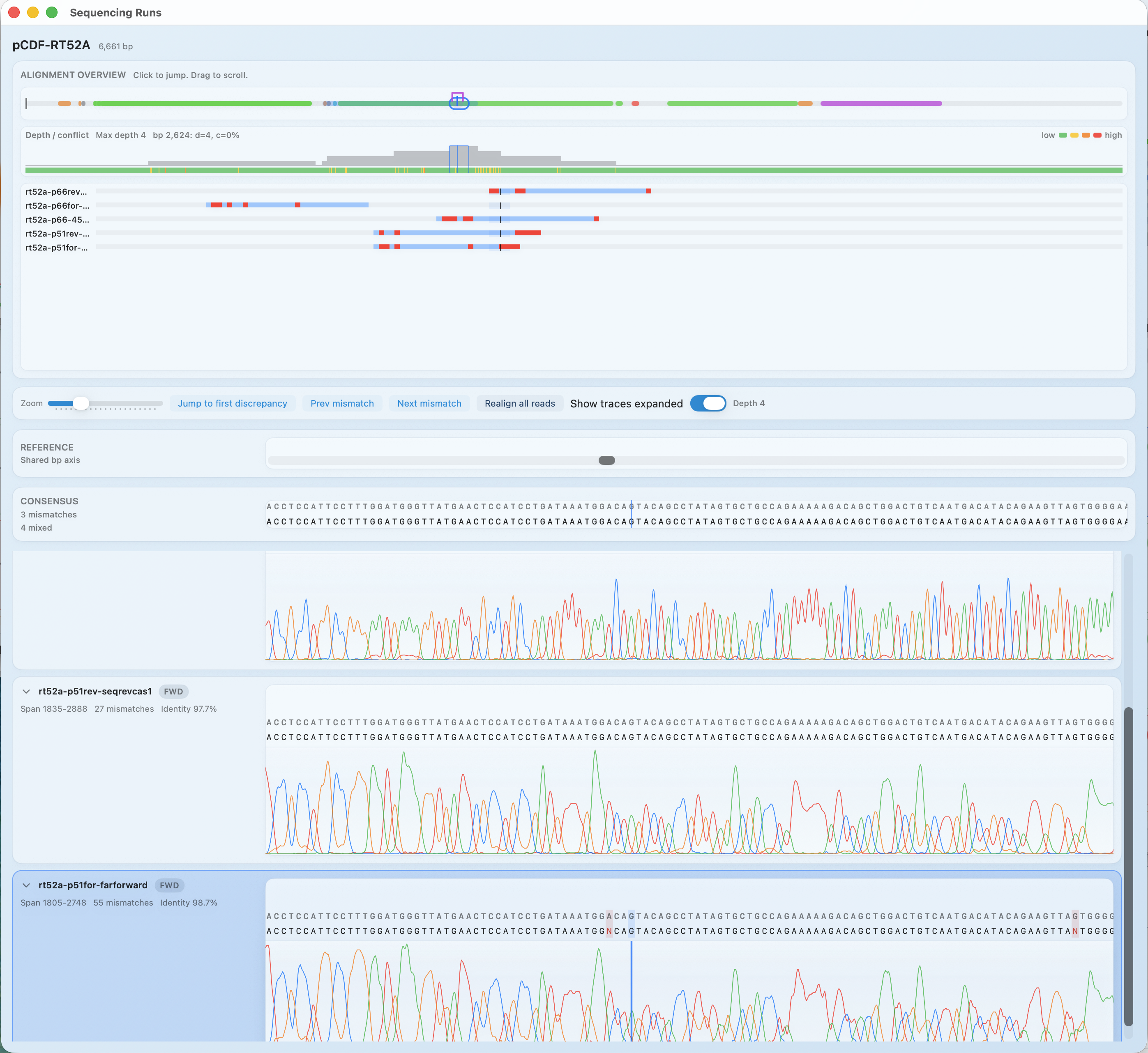
Sanger Validation
Open AB1 files and decide from real evidence
Use RayCrest as an AB1 viewer and chromatogram viewer for construct validation, so you can see mismatches where they occur instead of guessing from summaries.
Chromatogram review plus construct-linked QC notes and FASTQ support.
Multiple Sequence Alignment
Rerun protein MSA and keep the same result
AlignCove, RayCrest's built-in protein alignment tool, gives the same protein MSA result for the same input — making reruns and methods writeups easier to trust.
Built-in viewer, tree view, diagnostics, and repeatable outputs.
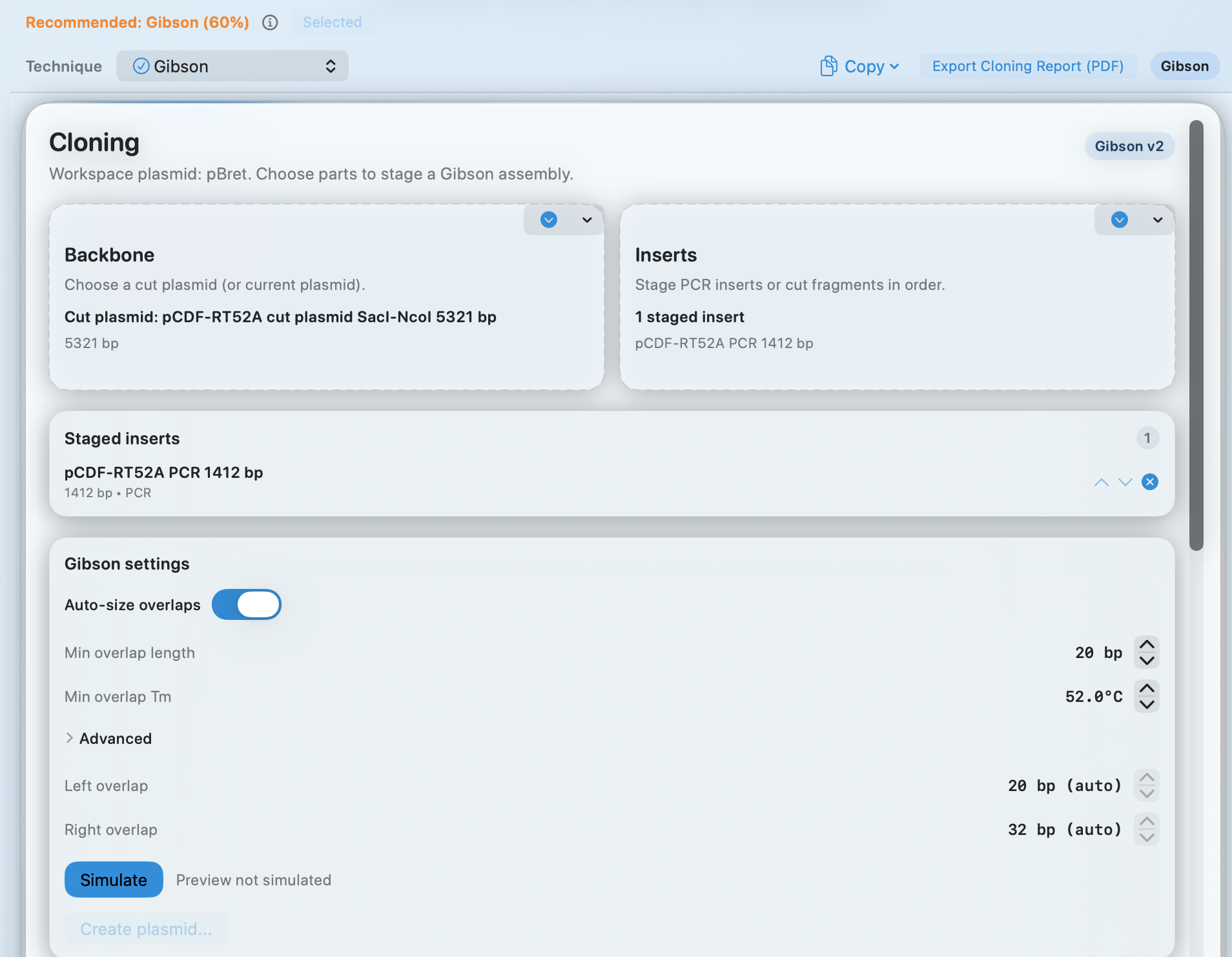
SnapGene Interoperability
Open SnapGene .dna files and keep working locally
Bring verified file types into the same Mac workflow you use for editing, validation, and map export.
Verified inputs: GenBank, FASTA, SnapGene .dna, Sanger .ab1/.abi, and FASTQ.
PROTEIN DESIGN
Codon optimization and reverse translation tools stay close to construct work
Use codon optimization and reverse translation tools without leaving the construct workflow.
Codon Optimization
Reverse translate and optimize coding sequences without leaving the construct workspace
Optimize codon usage while preserving the encoded amino-acid sequence, then review the result alongside your plasmid and primer work.
Current UI modes include Classic and Harmonize.
Reverse Translate accepts raw amino-acid text or one protein FASTA record.
PRODUCT SURFACES
Where those workflows live in the product
RayCrest centers the workflow around three main surfaces: construct work, evidence review,
and figure-ready output.
Sequence Workspace
Alignments + QC
Presentation Quality Figures
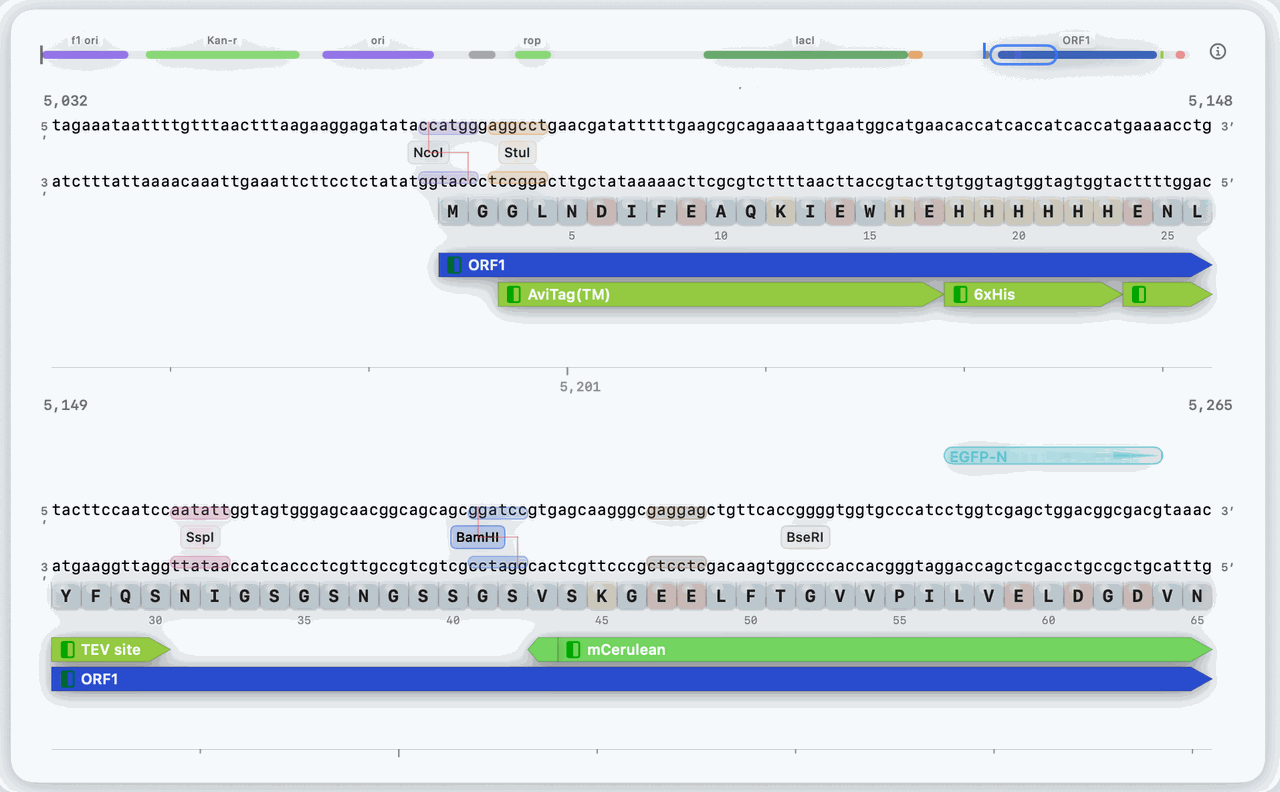
Sequence Workspace
Plasmid editing, primer design, and map-aware construct work all live in one workspace.
Edit constructs with annotations and overlays in place
Launch primer, digest, and cloning workflows from the active sequence
See Sequence Workspace
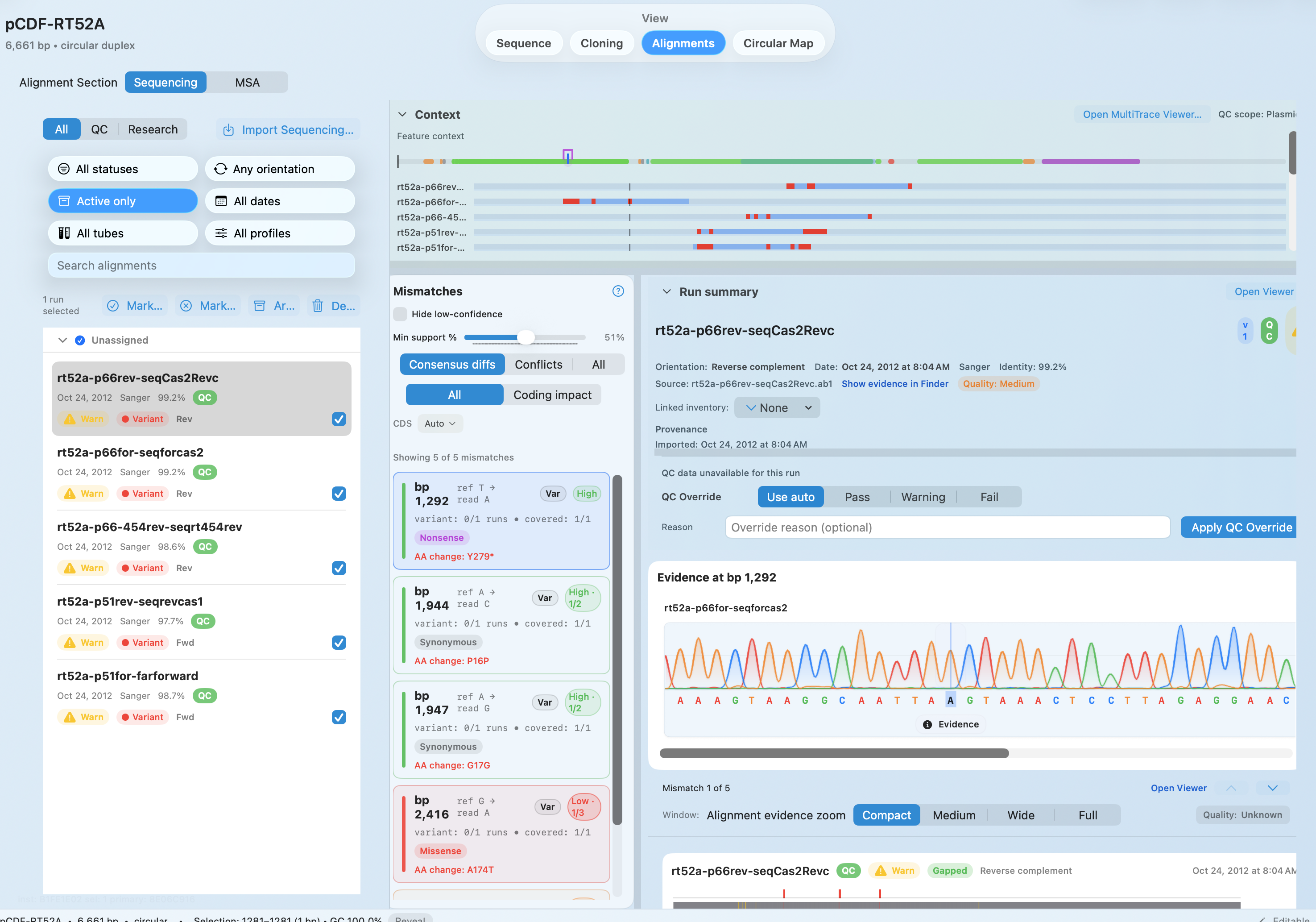
Alignments + QC
AB1 and FASTQ-backed validation plus repeatable protein MSA live in one review surface.
Jump from mismatches to chromatogram or FASTQ evidence
Keep QC decisions and AlignCove runs tied to the construct
See Alignments + QC
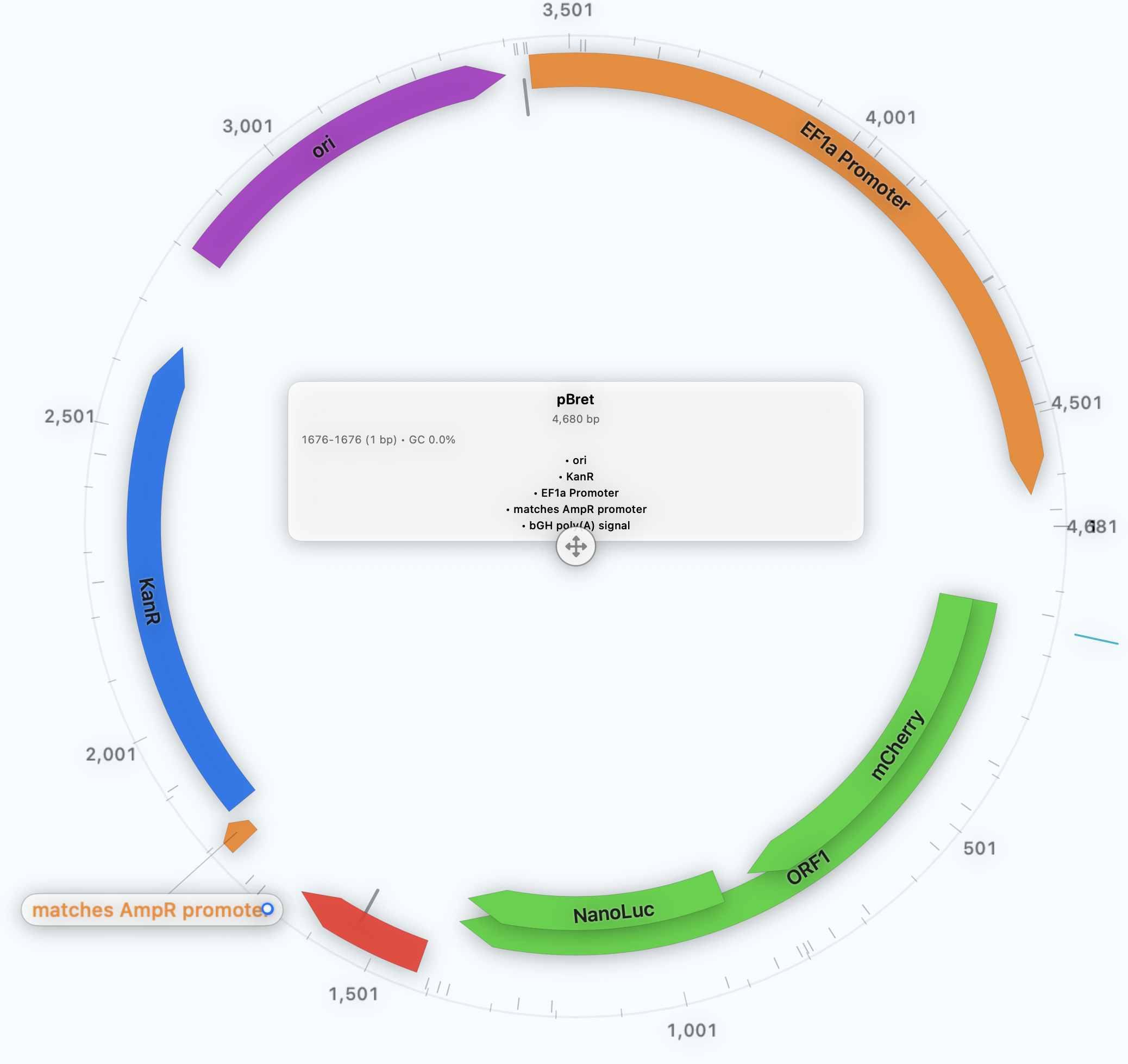
Presentation Figures
Publication-ready plasmid maps and vector-style outputs come from the same reviewed construct.
Circular, linear, and rectangular outputs from annotated constructs
Reusable styles for slides, manuscripts, and collaborator handoffs
See Presentation Figures
REAL CONSTRAINTS
Built for real biological constraints
RayCrest is built for the parts of bench work that become hard to trust when editing,
evidence, and outputs live in separate tools.
Deterministic protein alignment
Same input, same protein MSA output.
Constraint-aware codon optimization
Codon optimization surfaces CAI, GC %, CpG, repeats, and motif burden in one reviewable run.
Evidence-linked sequencing review
Jump from mismatch calls to chromatogram and FASTQ evidence instead of stopping at summaries.
Map output tied to the construct
Export plasmid maps from the same reviewed sequence rather than rebuilding them later.
CONNECTED WORKFLOW
From construct edit to validated handoff
RayCrest keeps design, simulation, validation, and history in one connected workflow so the
next step is always close to the construct.
Design
Edit the construct, keep annotations in view, and launch primer work from the active sequence.
Simulate
Check digest plans, virtual gels, and assembly logic before ordering or cutting.
Validate
Confirm mismatches with Sanger chromatograms or FASTQ support at the bases that matter.
Track
Keep runs, decisions, and construct history together so handoffs and writeups stay clear.
WHY MACOS-NATIVE
Local-first on Mac, on purpose
RayCrest is built for local files, native Mac behavior, and bench work that can't afford
browser delays or network dependencies.
Fast on everyday bench tasks
Edit, review, and export in a native Mac app that stays quick on everyday tasks.
Local files stay under your control
Work with local files and familiar privacy controls even when Wi-Fi or VPN gets in the way.
Less friction between tasks
Use familiar windows, shortcuts, and file behavior instead of bouncing across browser tabs.
ACCESS + BACKGROUND
Simple access and a brief background
Licensing
MDS will launch as an inexpensive individual subscription on the Mac App Store.
Invite-only beta access is available now for active bench workflows.
View licensing details
About MDS
MDS started as a practical workspace for molecular cloning and protein engineering, built
to keep design, evidence, and shareable outputs attached to the same construct.
Try RayCrest on your own bench workflow
Request beta access and tell us whether plasmid editing, primer design, sequencing validation, MSA, or map export matters most to your team.