Rerun the same input and keep the same result
Stable outputs are easier to compare across experiments, writeups, and review cycles.
MULTIPLE SEQUENCE ALIGNMENT
RayCrest MDS uses AlignCove for protein multiple sequence alignment, so the same input produces the same output on every rerun. The alignment viewer, tree, and run diagnostics all stay in the same workflow — no exporting to a separate tool.
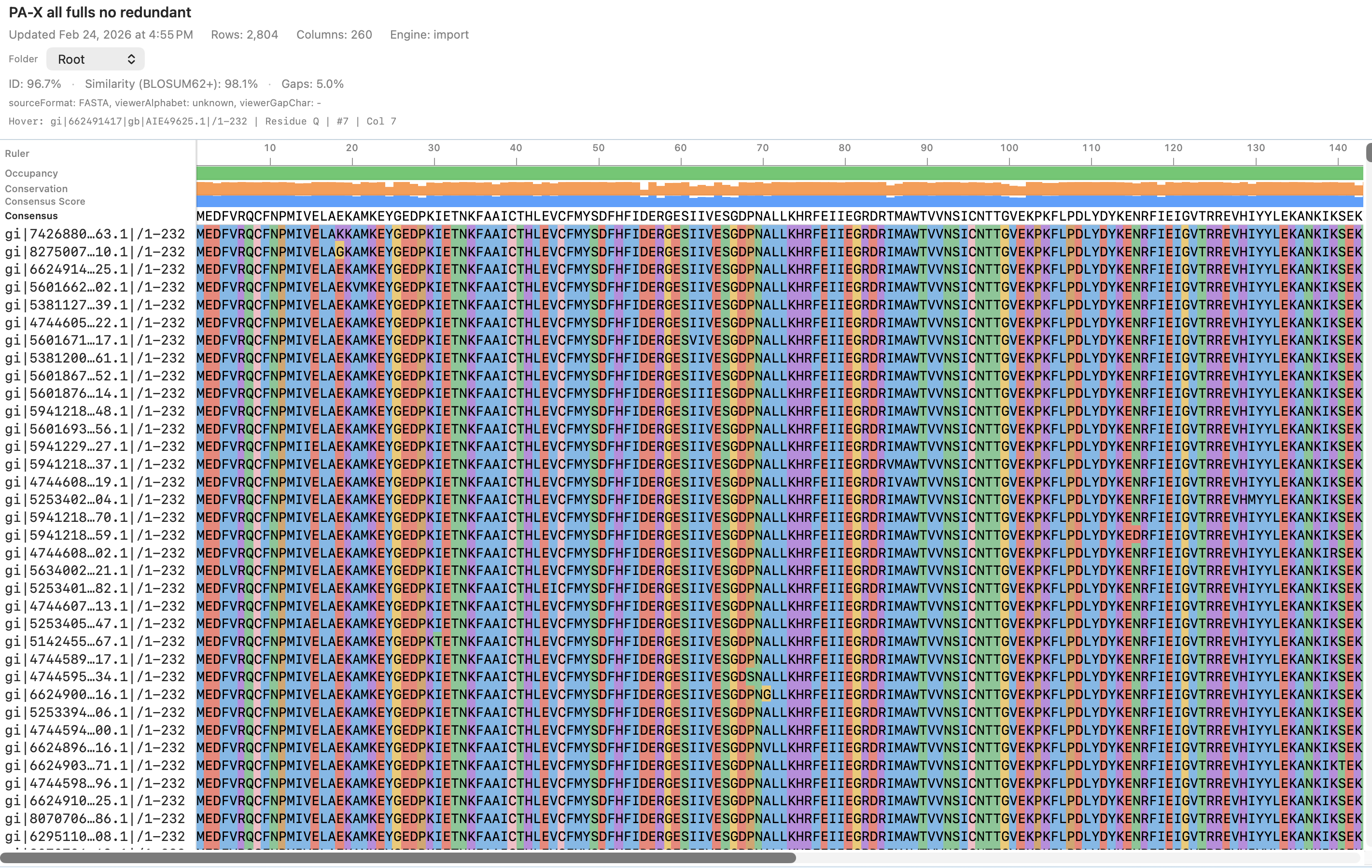
OUTCOMES
Stable outputs are easier to compare across experiments, writeups, and review cycles.
Protein alignment, plasmid editing, and validation all live in the same Mac app — no exporting to a separate tool.
Use viewer and tree outputs in the same workflow when you need to explain the result.
When it's time to write methods or review results, the diagnostics are right there with the alignment — not in a detached folder.
FEATURE WALKTHROUGH
A good alignment workflow isn't just about the first run — it's about whether reruns still support the same comparison and methods story.
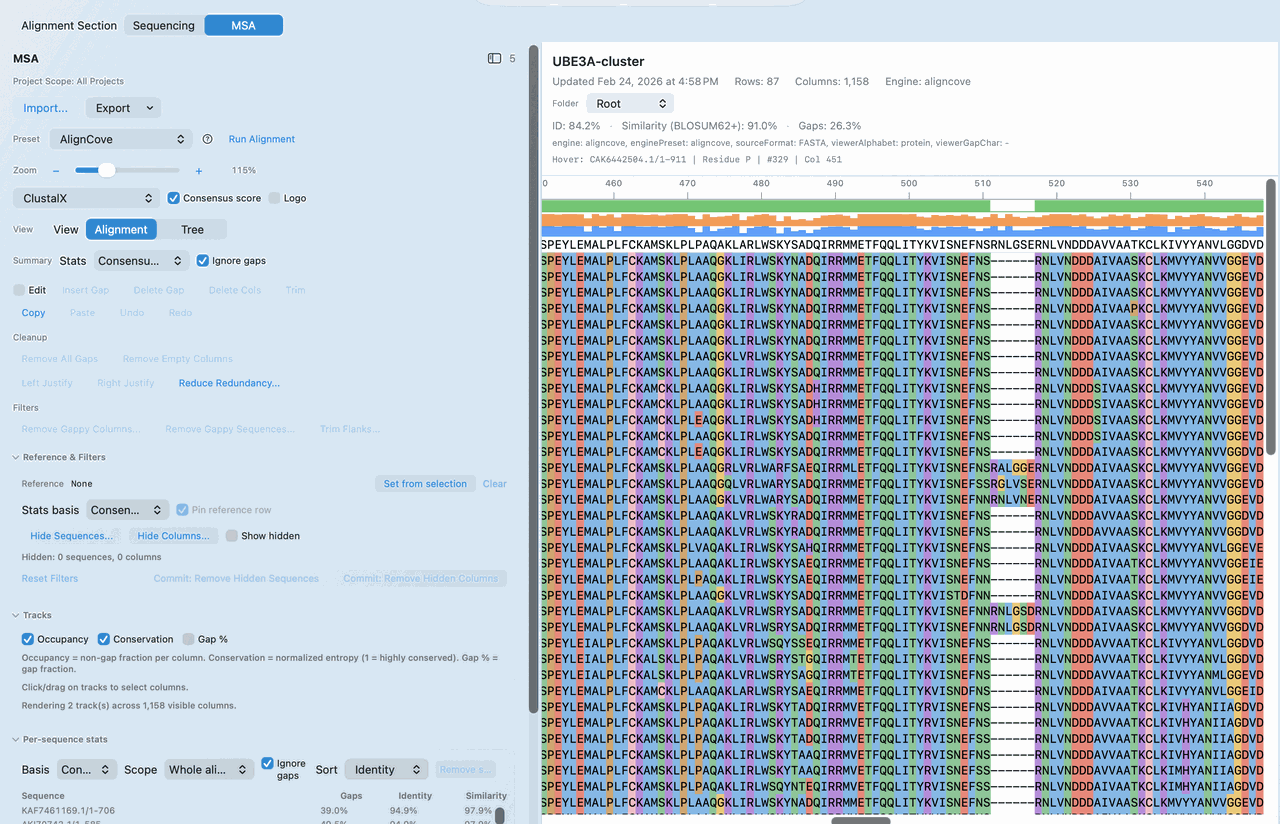
AlignCove is RayCrest's built-in protein alignment engine for teams that care about repeatable outputs and a workflow they can explain later.
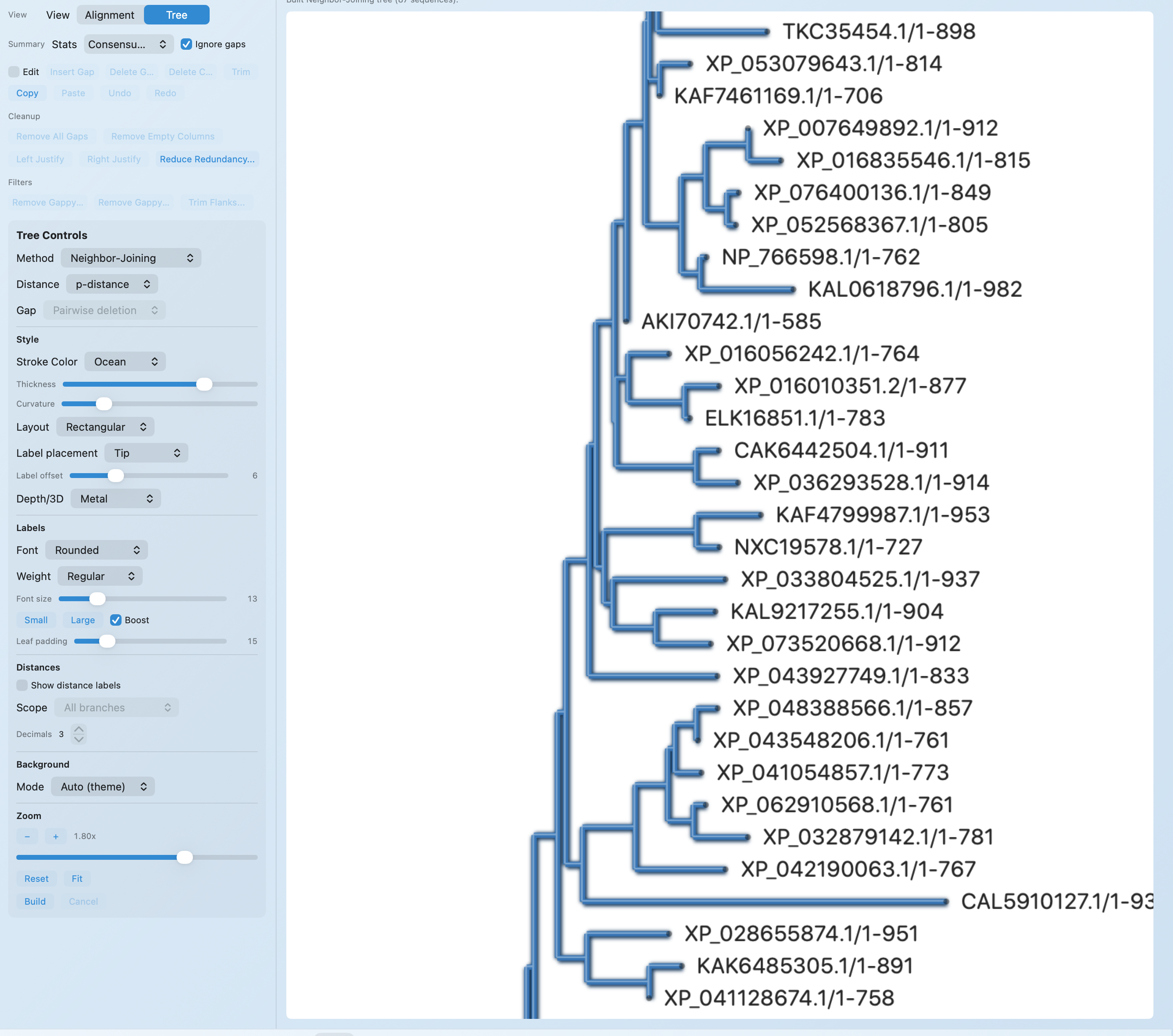
After the run, RayCrest keeps the alignment viewer, tree view, and diagnostics close to the result so you can inspect or explain it without another export step.
TECHNICAL DETAILS
AlignCove is positioned around repeatability, so reruns can be compared directly instead of treated as a moving target.
WORKFLOW + OUTPUTS
Choose the protein inputs and start an AlignCove run from the alignment workflow.
Review the alignment result, diagnostics, and tree output without leaving the app.
Rerun later with the same input and keep a comparable result for review or methods support.
RELATED PAGES
Validation
Use the broader validation workflow when the next question is construct evidence, not family-wide comparison.
AB1 Viewer
Keep chromatogram review and construct-linked QC decisions close to the same app and workflow.
Plasmid Work
RayCrest is strongest when alignment work sits alongside the plasmid and validation workflows that need the result.
Protein Design
When an alignment result drives a coding-sequence decision, continue the protein-to-DNA design loop in the same app.
Access
Tell us if AlignCove, deterministic reruns, or keeping protein alignment near construct work is what your team needs most.
Request beta access for a protein alignment workflow that stays stable across reruns and connected to your plasmid, validation, and primer work.