01
Work locally when the network is in the way
Design, review, and export on files stored on your Mac so VPN hiccups or cloud sync do not stall active bench work.
WHY RAYCREST MDS
RayCrest is built for bench teams that want plasmid editing, primer design, sequencing validation, repeatable protein MSA, and publication-ready map export in one local-first macOS workflow.
WHAT MAKES MDS DIFFERENT
01
Design, review, and export on files stored on your Mac so VPN hiccups or cloud sync do not stall active bench work.
02
Inspect Sanger or FASTQ evidence at the bases behind each call, with QC history tied to the construct.
03
AlignCove, the built-in protein alignment engine, produces consistent MSA output across repeated runs.
04
Generate clean circular, linear, and rectangular maps straight from the construct you already reviewed.
05
Start from common bench file formats and keep working in the same native Mac workflow instead of handing off between utilities.
KEY FEATURES
These are the parts of MDS most bench teams reach for during design, sequencing review, and figure prep.
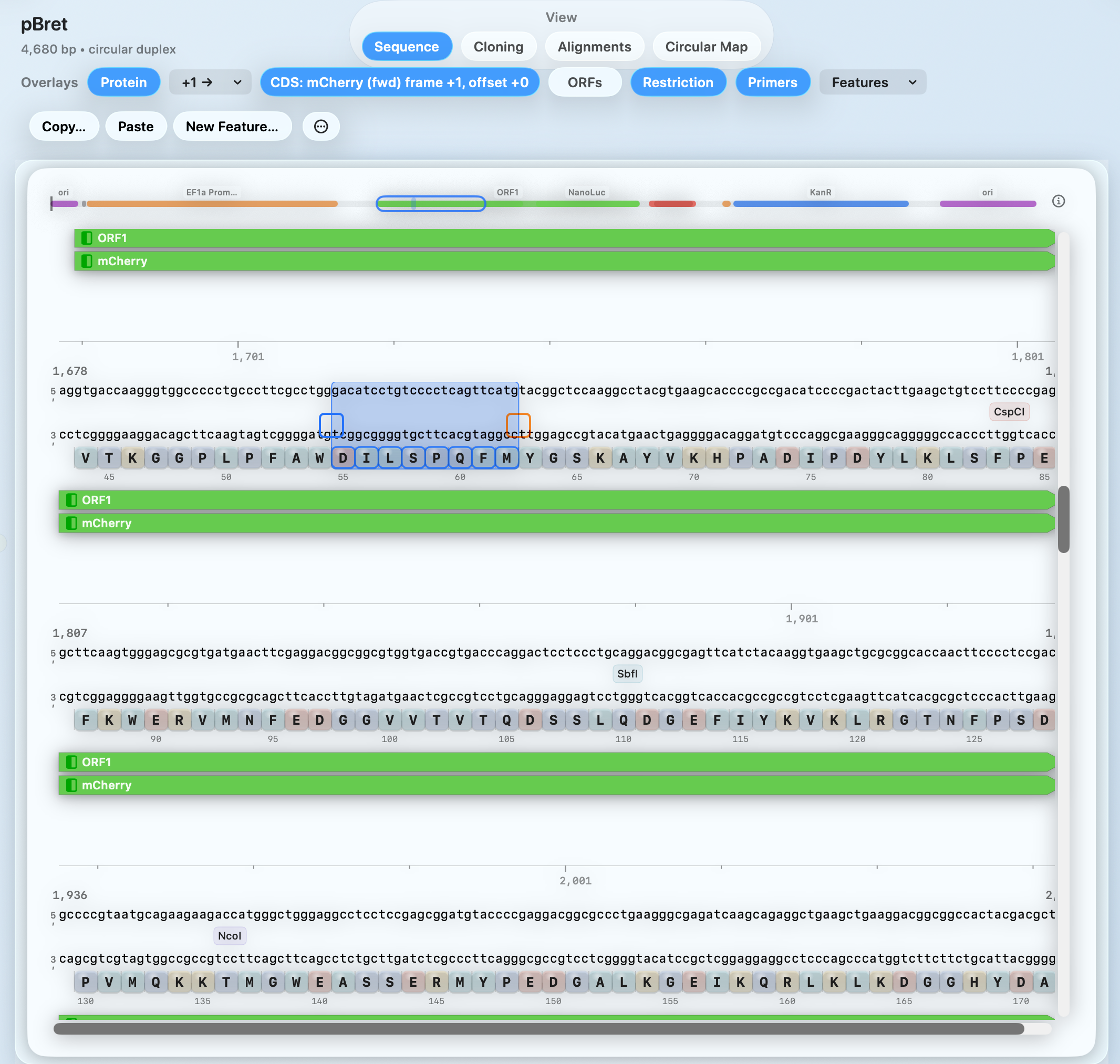
PRIMARY WORKSPACE
Use one plasmid editor and plasmid map maker for sequence changes, map context, primer design, and digest planning.
See Sequence Workspace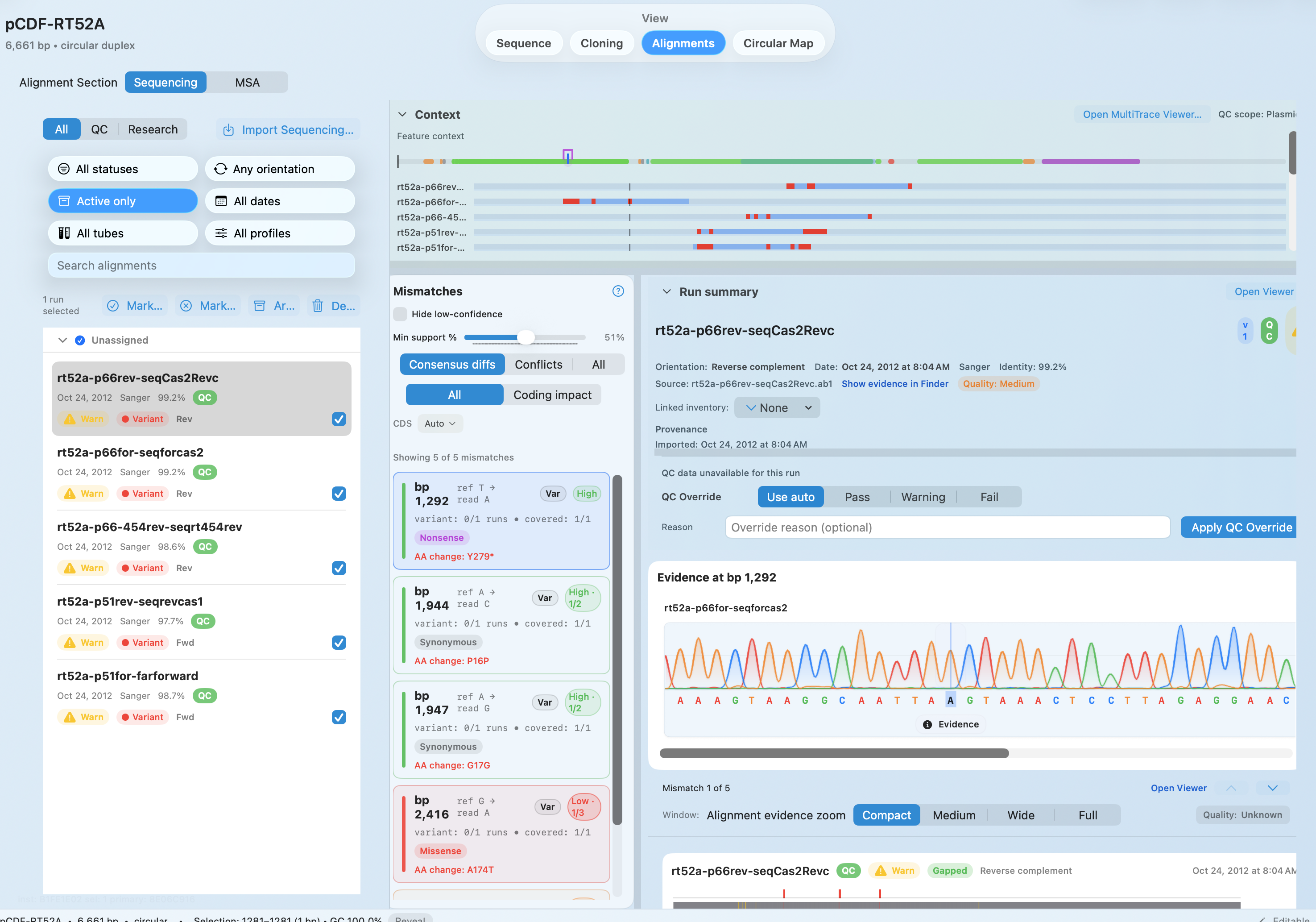
QC + VALIDATION
Review AB1 traces, FASTQ evidence, and repeatable protein MSA results before moving constructs forward.
See Alignments + QC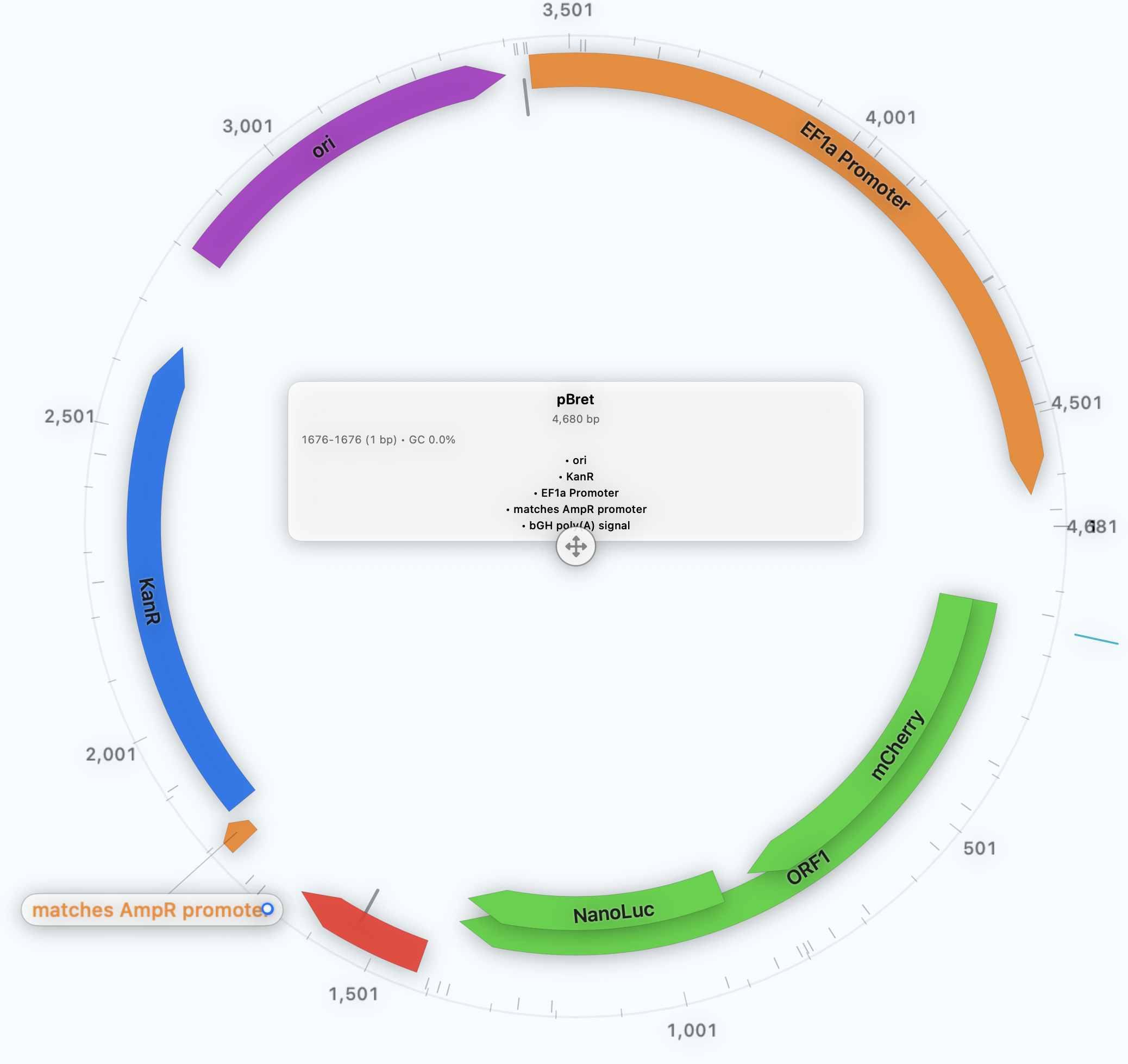
COMMUNICATION
Export publication-ready plasmid maps with reusable styling from the same construct and validation context.
See Presentation Quality FiguresBUYER SUMMARY
RayCrest is strongest when the same team needs plasmid editing, primer design, AB1 and FASTQ review, repeatable protein MSA, and coding-sequence design — all without rebuilding context in separate tools.
Plasmid Editing
Sequence edits, annotations, overlays, and map output stay tied to the same construct record.
Primer Design
Primer Wizard, digest planning, and cloning workflows live next to the sequence they affect.
Validation
Trace mismatch calls to chromatogram and read-level evidence before approving construct outcomes.
MSA
AlignCove, RayCrest's built-in sequence alignment software, gives teams the same protein MSA output on reruns — making methods writeups and result comparisons more reliable.
Protein Design
Reverse translate protein input to coding DNA, optimize coding regions while preserving amino-acid sequence, and keep the result close to AlignCove and construct editing.
Interoperability
Bring common sequence and validation files into a local-first workflow built for editing, validation, and map export.
IN PRACTICE
The change is usually less reconstruction between steps: fewer places where the same construct has to be redrawn, re-explained, or revalidated.
MAPS
The construct you edited is already the construct you export, so figure prep does not start with redrawing.
EVIDENCE
Jump from a flagged mismatch to chromatogram and FASTQ evidence when the call matters.
CONTEXT
Primer choices, QC notes, and map output stay attached to the same sequence record.
PROTEIN
Move from protein sequence to reverse-translated coding DNA inside the same construct workflow.
WHERE MDS FITS
MDS is strongest when the same team needs to design constructs, review sequencing evidence, and share clean outputs without rebuilding context.
You'll love MDS if most days start with plasmid maps, primer plans, digests, and assembly choices.
You'll love MDS if you regularly review Sanger or FASTQ results before moving a construct forward.
You'll love MDS if you frequently make maps for ordering, meetings, or manuscripts.
INTEROPERABILITY
RayCrest imports verified file types you already use: GenBank, FASTA, SnapGene .dna, Sanger .ab1/.abi, and FASTQ — all in a local-first macOS workflow on your own machine.
Sequence Files
Import GenBank files into the same construct workspace used for editing, validation, and figure export.
Sequence Files
Start from FASTA when you need a simple sequence path before annotations, primers, or review workflows.
SnapGene
.dna importImport compatible SnapGene files and continue work in the local Mac workflow.
Sanger
.ab1 / .abiOpen verified trace formats for evidence-linked mismatch review and chromatogram inspection.
FASTQ
Bring FASTQ runs into the same QC workflow for support-depth review and construct-linked decisions.
Local Workflow
Keep files, review history, and exports on your Mac when bench work cannot wait for the network.
REFERENCE MATRIX
Full feature comparison across MDS and common alternatives.
| Software | Plasmid map; edit/annotate | Auto‑annotate (feature DB) | Restriction analysis & virtual digest/gel | Primer design & in‑silico PCR | Site‑directed mutagenesis | Cloning simulation (Restriction / Gibson / Golden Gate / In‑Fusion‑NEBuilder / Gateway / TA‑TOPO) | Sanger traces & assembly | Multiple sequence alignment | CRISPR gRNA design/analysis | Codon optimization / reverse translate | BLAST | NGS analysis | ELN / LIMS / Registry | Notes |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MDS | + | + (Library learns your annotations) | + (Restriction Analyzer + gel sim) | + (Primer Wizard; in‑silico PCR) | + | + / + / + / + / + / + | + / + | + AlignCove Mac native alignment | - | + (reverse translate; codon optimization in Classic/Harmonize modes) | + (NCBI + local/custom) | - | Registry | Tracks cloning history, curated plasmid and primer library, local storage and calculations |
| SnapGene | + | + (common feature DB) | + (incl. gel sim) | + (auto primer design; in‑silico PCR) | + | + / + / + / + / + / + (incl. multisite Gateway; TA/GC/TOPO) | + / + (view .ab1; CAP3 contigs) | + (ClustalΩ/MAFFT/MUSCLE/T‑Coffee) | - (no dedicated CRISPR tools) | + (reverse translate + codon usage tools) | + (NCBI BLAST via Tools menu) | - | - | Also tracks cloning history, curated plasmid library. |
| Benchling (Molecular Biology) | + | + (feature libraries; bulk auto‑annotate) | + (virtual digest + predicted gel) | + (Primer Wizard; in‑silico PCR) | Partial (via primer workflows) | + / + / + / + (Homology/HiFi) / Partial (Gateway via homology; att sites required) / - | + (Sanger reads + consensus alignment) | + (DNA & AA MSA) | + (CRISPR guide design tool) | + (codon optimization & back‑translate) | + (BLASTn/BLASTp) | - (no native pipelines; integrates; don’t upload raw FASTQ) | + (ELN, Registry, LIMS) | Strong collaboration, APIs/SDK; antibody numbering & CDR annotations. Assembly tools support digest+ligate, Gibson, Golden Gate, and Homology; homology can model In‑Fusion/HiFi and some Gateway cases. |
| Geneious Prime | + | + | + | + | + | + / + / + / + / + / + | + / + | + | + (find sites, off‑targets, analyze edits) | + | + (NCBI + local/custom) | + (mapping, de novo, RNA‑Seq, variants via modules) | - | Workflows, plugins, GenBank submission, CLI/API. |
| DNASTAR Lasergene (SeqBuilder Pro, etc.) | + | + | + | + | + | + / + / + / + / + / + (incl. LIC/SLIC/CPEC/SLiCE; MultiSite Gateway Pro) | + / + | + | - (no dedicated gRNA design tool evident) | Partial (translation/back‑translation) | + (NCBI integration; BLAST) | + (Lasergene Genomics suite) | - | Broad file import (incl. SnapGene/Geneious); virtual cloning workflows. |
| ApE (A Plasmid Editor) | + (circular/linear) | + (custom feature libs) | + (Dam/Dcm aware); virtual digest | + (primer find + in‑silico PCR tool) | Basic | + / + / + / + (Gibson/HiFi/InFusion) / + (Gateway/recombination) / Partial (TA; TOPO not evident) | Partial (ABI trace view + align to reference) | Basic (pairwise) | Partial (sgRNA analysis tool) | Partial (reverse translate; no codon optimization) | + (NCBI/Wormbase BLAST) | - | - | Free, cross‑platform; vector graphics export. Also includes BLAST, Gateway recombination assembler, and sgRNA analysis. |
Design, validate, and export without rebuilding context across separate tools.
See the Sequence Workspace