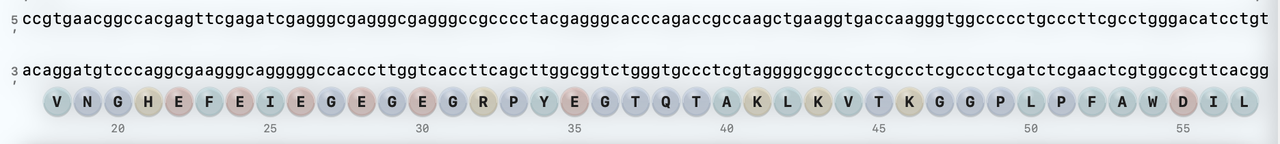
Paste amino-acid sequence or load a FASTA record.
CODON OPTIMIZATION + REVERSE TRANSLATE
Codon optimization tool and reverse translation, built into your Mac construct workflow
Use the reverse translation tool to reverse translate protein sequences to coding DNA and optimize in-frame coding targets while preserving the encoded amino-acid sequence. Everything stays in the same local-first macOS workflow as your sequence editing, plasmid maps, primer design, and AlignCove runs — no context switching required.
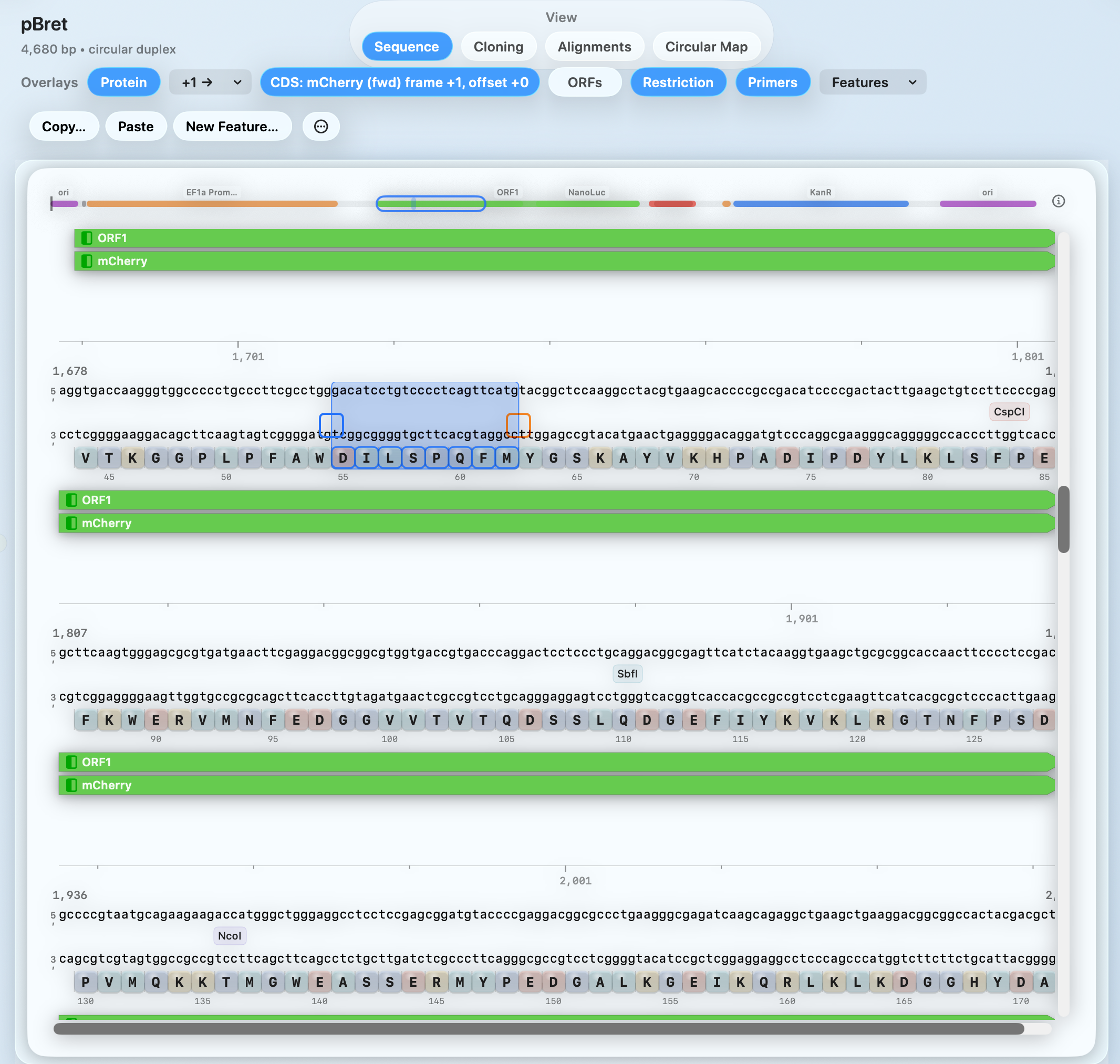
VISUAL WORKFLOW
From protein sequence to optimized coding DNA
Reverse translate protein sequences to coding DNA while preserving the protein sequence.

Run Classic or Harmonize optimization modes.
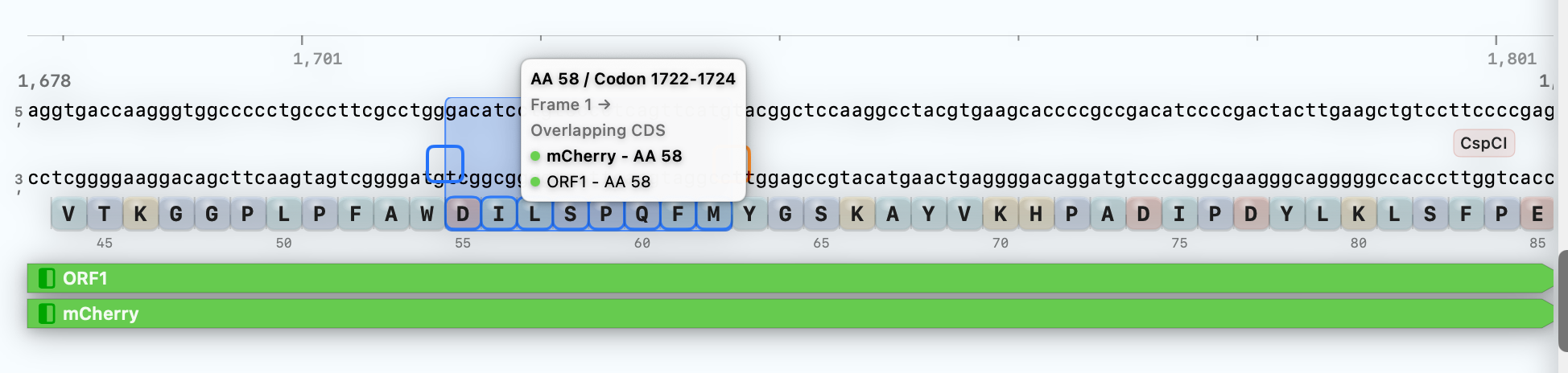
Use the optimized DNA directly in plasmid editing, primer design, and map export.
OPTIMIZATION MODES
Choose the optimization mode that fits your protein
Both modes use a deterministic optimizer — the same coding target and settings always produce the same optimized sequence and metrics, so results are reproducible and reviewable.
Classic Optimization
Maximize expression in your target organism
Use Classic when you want to improve codon efficiency and sequence
quality in one pass without changing the encoded amino-acid sequence.
- Increases codon adaptation index (CAI) while preserving the amino-acid sequence.
- Maintains codon diversity and avoids repetitive DNA where possible.
- Minimizes problematic motifs and keeps GC and CpG levels within safe ranges.
Best for: gene synthesis, recombinant protein expression, and plasmid construct design.
Harmonize Optimization
Match the original organism's translation rhythm
Use Harmonize when re-expressing a protein from another organism and
folding kinetics matter. It adapts the CDS for the target host while preserving
relative codon-usage patterns from the source.
- Compares codon rarity between source and target organisms.
- Maintains relative codon-usage patterns instead of only favoring frequent codons.
- Preserves translation pauses that may matter for protein folding.
Best for: complex proteins, multi-domain proteins, and cases where folding kinetics matter.
BIOLOGICAL CONSTRAINTS
Sequence properties evaluated on every run
The optimizer evaluates several biological sequence properties:
- Codon Adaptation Index (CAI)
- GC content windows
- CpG frequency
- Repeated DNA motifs
- Transcription and translation motif patterns
- Out-of-frame start codons
RESULT PROOF
What an optimization run reports
RayCrest reports before-and-after optimization metrics so the result can be reviewed, not treated as a black box. This example surfaces CAI, GC %, CpG count, repeat burden, motif burden, and deterministic output in one place before you accept the coding-sequence change.
Optimization Report
Review the optimization result before accepting the coding-sequence change
- Mode:
Classic - Target: selection of 1,674 nt (558 codons)
- Codon changes: 282 mutations in this example
- Output style: deterministic for the same input and settings
Before / after metrics
Optimization summary from one Classic result
| Metric | Before | After |
|---|---|---|
| CAI (human_nuclear) | 0.6501 | 0.8850 |
| %GC | 37.9% | 53.9% |
| CpG count | 1 | 15 |
| Repeat 8-mers | 82 | 5 |
| Motif hits (PAS / splice-like) | 17 / 29 | 2 / 26 |
Example values above are shown to demonstrate the kind of before/after report RayCrest exposes. CAI, GC %, CpG count, repeat burden, motif burden, and deterministic output stay reviewable in one result view instead of collapsing into a hidden single-number decision.
UI PROOF
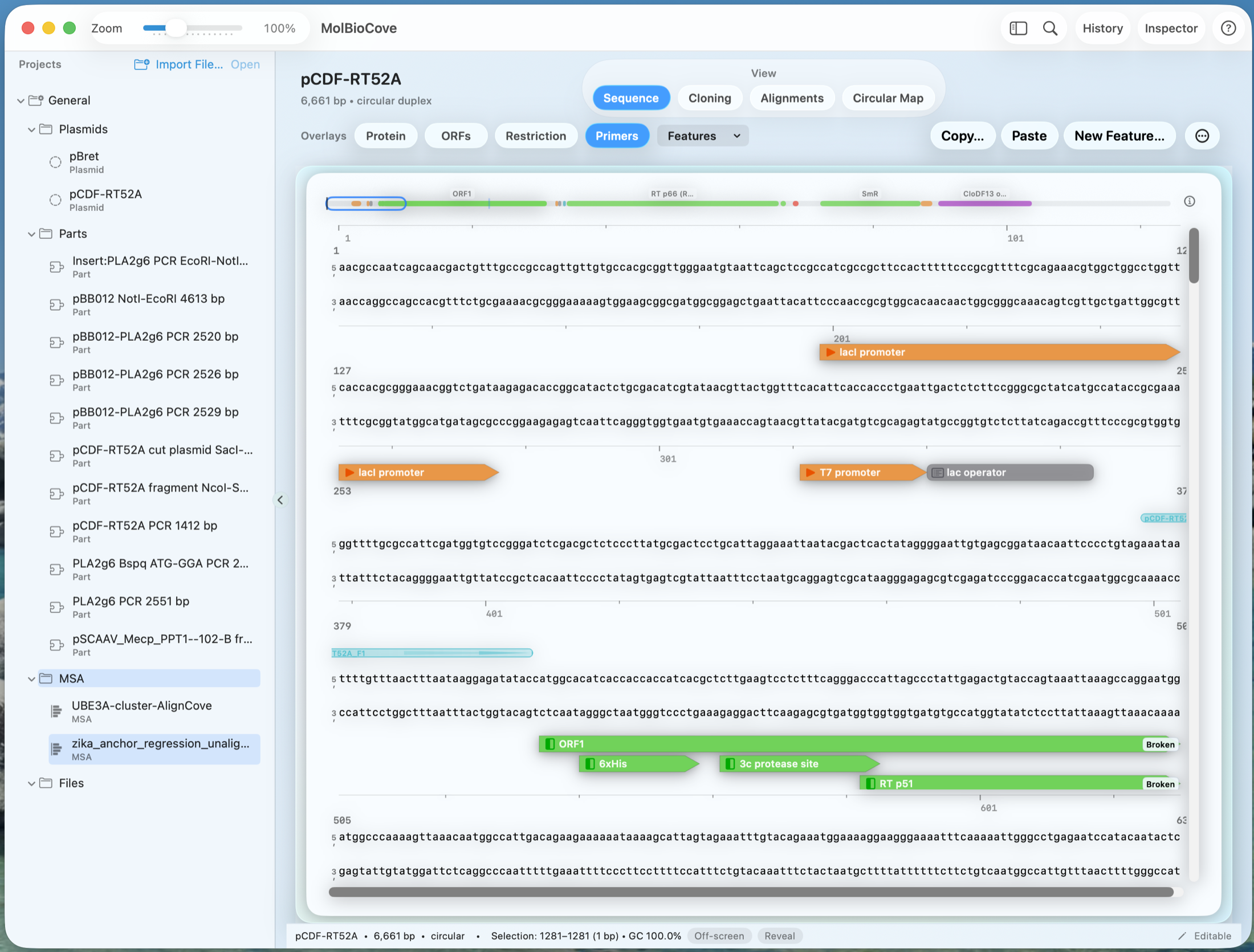
See the workflow in the interface

Keep coding-sequence design in the same construct workspace.
Reverse-translated or optimized DNA stays alongside sequence, protein, and annotation context — no separate utility needed.

Review codon and amino-acid indexing inline, alongside the in-frame targets you're working with.
Codon and amino-acid context stays visible right where the optimized sequence continues downstream.
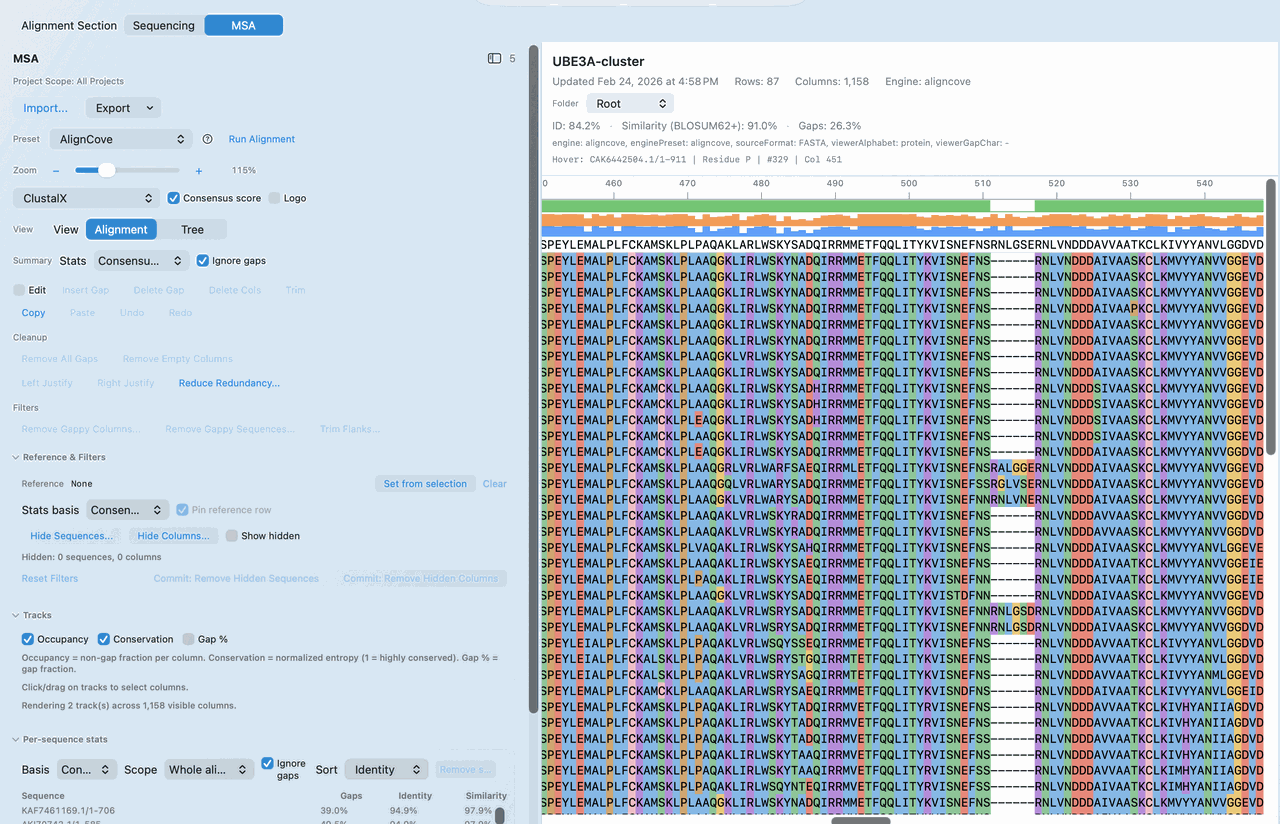
Optional: start from AlignCove when protein comparison informs the next coding-sequence decision.
Run a protein MSA first, then carry that result directly into reverse translate or codon optimization.
CURRENT SCOPE
What RayCrest codon optimization supports today
| Supported now | Not claimed |
|---|---|
|
|
CONNECTED WORKFLOW
Keep codon work connected to the rest of RayCrest
Each optimized sequence flows directly into the construct, alignment, and primer surfaces you're already working in.
Sequence Workspace
Save the optimized DNA into the same construct record
Go straight back to plasmid editing, annotation review, and map-aware construct work without rebuilding context.
MSA
Use AlignCove when protein comparison is the entry point
Start from protein MSA when comparison should drive the next codon or reverse-translate decision.
Primer Design
Move into primer planning after the coding sequence changes
Open Primer Wizard when the next step is sequencing confirmation, cloning, or mutagenesis work on the updated construct.
Design coding sequences without leaving your construct workflow
Request beta access to keep protein-to-DNA design in the same local workflow as plasmid editing, MSA, validation, and map output.